
Get ready for Friday's eclipse!

A solar eclipse observed in Australia in 2012. Image: Davidfntau.
On Friday March 20, between around 9:25 GMT and 10:41 GMT, people in the UK will be able to witness a partial eclipse of the Sun! It's the first one since the total eclipse in 1999 and there won't be another one until 2026.
A solar eclipse occurs when the Moon moves in front of the Sun, blotting parts of it out and casting its shadow on the Earth. Some parts on the Earth will see a total eclipse, in which the Moon completely blocks the Sun, giving a great view of its outer atmosphere called the corona.
On Friday only a narrow path a few hundred kilometres wide will see a total eclipse. The only two landmasses in that path are the Faroe Islands and the arctic archipelago of Svalbard. In the UK the Sun will only be partially obscured to varying degrees: Edinburgh will see 93% of the Sun blotted out, Lerwick in the Shetland Isles 97%, and London 85%. In London the eclipse will start at 8:25GMT, peak at 9:31GMT and end at 10:41. Times vary slightly in other parts of the UK, with things happening earlier the further West or South you are.
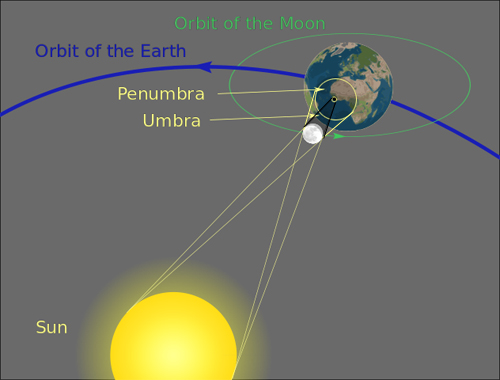
The geometry of an eclipse. In the region called the umbra, bounded by the small yellow circle, people see a total eclipse. In the penumbra, bounded by the larger yellow circle, people see a partial eclipse.
But even when the eclipse is only partial, there are still interesting things to look out for. You might see dark spots on the Sun — sunspots — which are caused by the magnetic field of the Sun dimming its light output. You might also be able to observe the outline of the Moon's surface, with its mountains and valleys. If you're somewhere where the sunlight passes through tree foliage, you'll see lots of images of the eclipse projected on the ground and other things around you, as the holes in the foliage act as pin hole cameras.

Pinhole images on a car during the 1998 eclipse in Guadeloupe.
It's really important that you don't look at the Sun directly, not even while it is being eclipsed, as this can seriously damage your eyes. Even sunglasses aren't good enough protection. But even if you haven't got specially produced eclipse viewing glasses, chances are there is something in your house that will help you view it. A colander and a piece of paper, for example. Standing with your back to the Sun, let the light pass through the colander onto the paper to see lots of images of the eclipsed Sun appear. You could also use a mirror to reflect the light onto a wall, or build your own pinhole camera. See this lovely booklet, produced by the Royal Astronomical Society, to find out more.
Now all we need is a clear sky!